2016 UCR Computational Chemistry & Biology Workshop
Thank you to all of the students who attended this year's workshop!
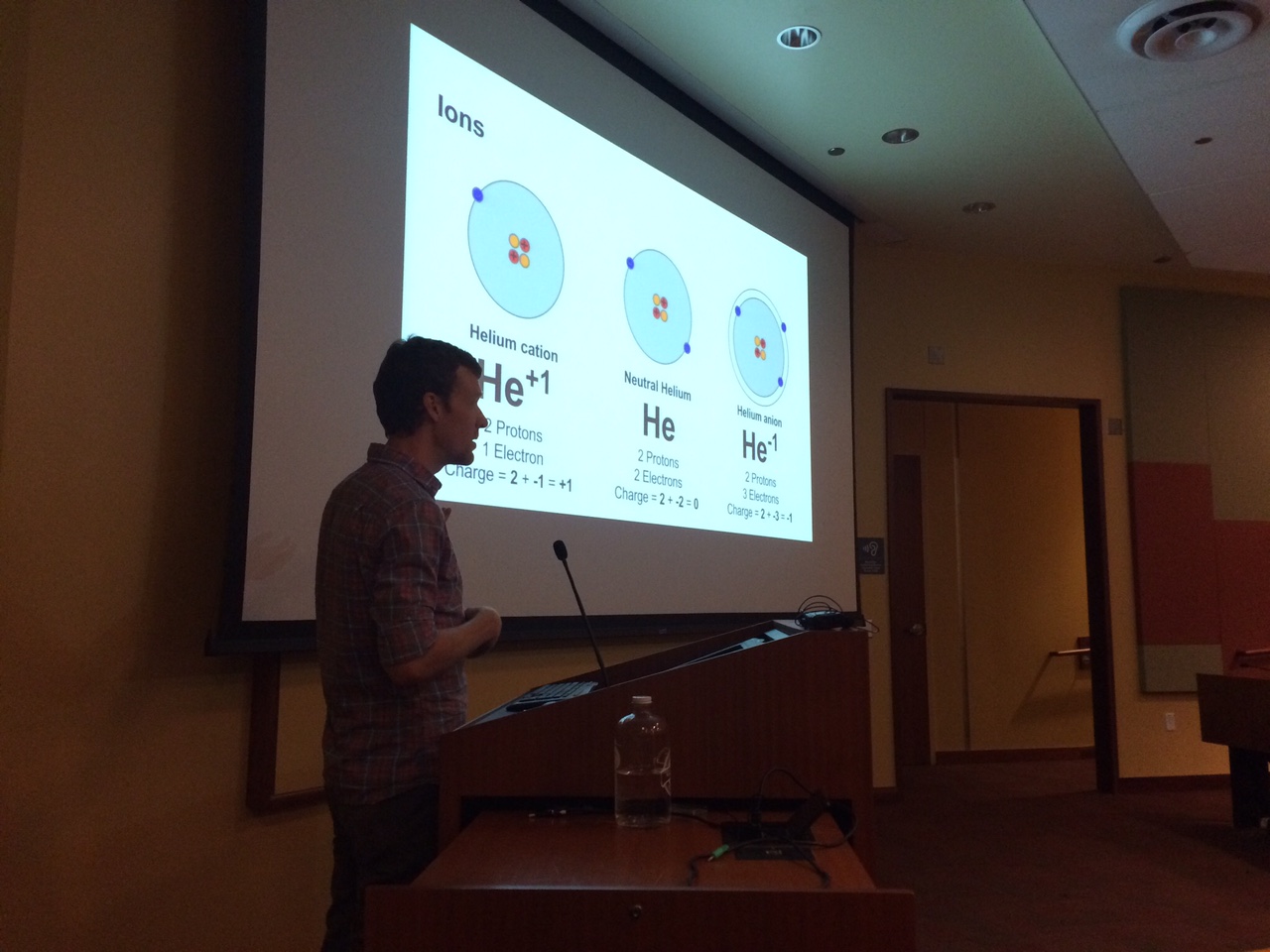

Co-organizer Dr. Christopher C. Roberts presenting basics of chemistry during the introductory lecture of the 2016 workshop.
 Participants of the 2016 workshop in the UCR Genomics Auditorium.
Participants of the 2016 workshop in the UCR Genomics Auditorium.
UCR Computational Chemistry & Biology Workshop
Recent advances in computers have provided powerful tools to study molecular binding, chemical catalysis, enzyme function, drug design/discovery, etc. In this workshop, meant for college students, we will introduce ideas about how we use computers to study biological systems. The workshop will cover both lectures and hands-on session, so please bring your laptop!
Itinerary
Day One - Tuesday August 2nd
| 9:00-10:00AM | Registration, Check-in, and Breakfast @ Genomics Building Lobby | |
| 10:00 AM | (Dr. Chang) | Welcome students, introduce group, etc. |
| (Dr. Roberts) | Overview of compuational methods, quick review of chemical concepts (atoms, molecules, bonding, forces), introduce Acetylcholine esterase system | |
| (Wanli You) | Molecular visualization with VMD and creation with Avogadro. | |
| 12:00 PM | Lunch Break | |
| 1:00 PM | Hands-on - Obtain experimental structures, visualize hydrogen bonds, ES field visualization, etc. |
Day Two - Wednesday August 3rd
| 9:00-10:00AM | Registration, Check-in, and Breakfast @ Genomics Building Lobby | |
| 10:00 AM | (Dr. Tang) | Basics of molecular mechanics |
| (Mark Raymundo) | Molecular Docking with AutoDock Vina | |
| Docking demonstration. | ||
| Hands-on session: Docking one of several acetylcholine esterase substrates or inhibitors. | ||
| 12:00 PM | Lunch Break | |
| 1:00 PM | (Dr. Tang) | Basics of dynamics |
| (Dr. Roberts) | Brownian dynamics: Problems it can solve, how it works, etc. | |
| Brief BD demo, show analysis of output | ||
| Hands-on session: Brownian dynamics analysis of Acetylcholine esterase + ligand/inhibitor |
Pre-Workshop Preparation
Participants will be learning the use of the Visual Molecular Dynamics (VMD) software available from the Theoretical and Compuational Biophysics Group at U. of Illinois at Urbana-Champaign. The software is available free of charge, and is available for Windows, Linux, and Mac OS X. Registration is required to download. Participants should download and install the software from the following links:
If you experience any difficulties with either downloading or installation, please contact Dr. Christopher C. Roberts.